A multiplex assay capable of detecting all known pathogens of concern in a single small blood sample with high sensitivity and specificity could significantly increase the safety of the blood supply. and the Procleix Ultrio Plus,Gen-Probe, Inc.) use PCR or transcription-mediated amplification methodology for multiplex detection of HBV, HCV, and HIV 1–2. Two FDA-approved blood donor screening assays (Cobas Taqscreen MPX Test, Roche Molecular Systems, Inc. Multiplex PCR-based devices for testing blood-borne pathogens are limited. Nearly all the other agents require individual qPCR or serologic testing and it is logistically impractical and cost prohibitive to test all known and potential agents individually. Only prions cannot be detected by currently available technology. The American Association of Blood Banks (AABB) Transfusion–Transmitted Diseases Committee produced a list of over 30 pathogens of concern for transmission via blood that included bacteria, parasites, prions and viruses. HTLV, syphilis and Chagas antibody testing fail to detect these pathogens during early infection and Chagas is screened only once on samples from first-time blood donors. Food and Drug Administration (FDA) licensed methods for infectious disease screening of donor blood include (1) NAT for Hepatitis B virus (HBV), Hepatitis C virus (HCV), HIV 1 and 2, Babesia, West Nile virus (WNV) and Zika virus (ZIKV) and (2) immunoassays for HBV, HCV, HIV-1, 2, cytomegalovirus (CMV), human T cell lymphotropic virus I and II (HTLV), Treponema pallidum (syphilis) and Trypanosoma cruzi (Chagas). Though screening of these blood units using serologic and nucleic acid testing (NAT) has greatly reduced the risk of some transfusion–transmitted infections, the vast majority of bloodborne agents are not screened. This microarray-based multi-pathogen screening platform accurately and reproducibly detected individual and mixed RNA viruses in one test from single samples with limits of detection as low as 10 2 copies mL.Įach year over 112 million blood donations are collected globally and nearly 21 million blood components are transfused in the U.S.

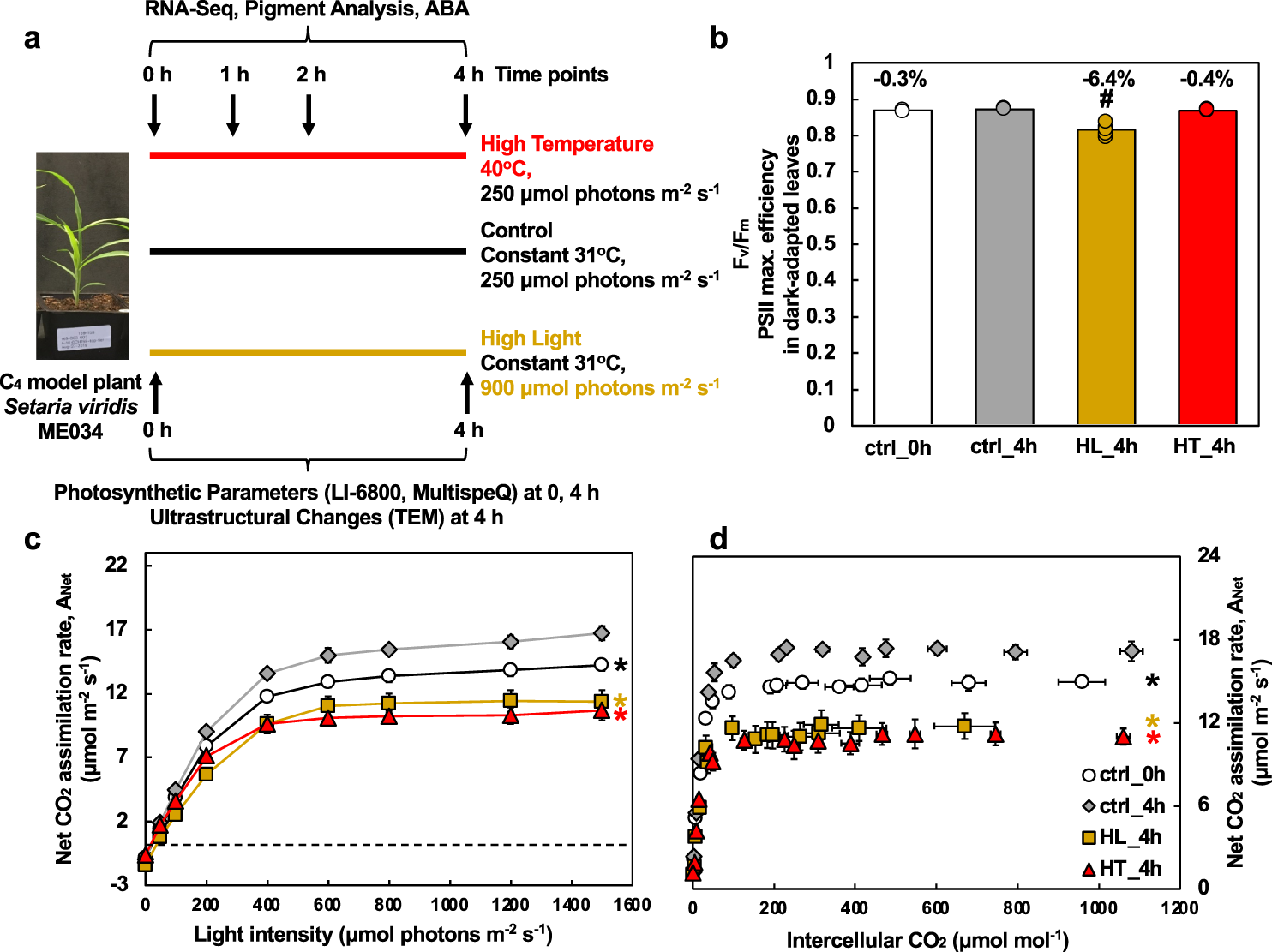
In mixtures of 3–8 different viruses, all were correctly identified between 10 5 and 10 3 copies/mL. Hepatitis A, B and C, Chikungunya, dengue 1–4, HIV 1–2, HTLV I–II, West Nile and Zika viruses were all correctly identified by the pathogen chip within the range of 10 5 to 10 2 copies/mL hepatitis E virus from 10 5 to 10 4. RNA-viruses detection and concentration were validated by RT-qPCR.

Differentially concentrated positive plasma samples were used to evaluate performance and limits of detection in the context of individual pathogens or combinations to simulate coinfection.
Viruses were targeted for detection by 1769 oligonucleotide probes selected by Agilent eArray software.
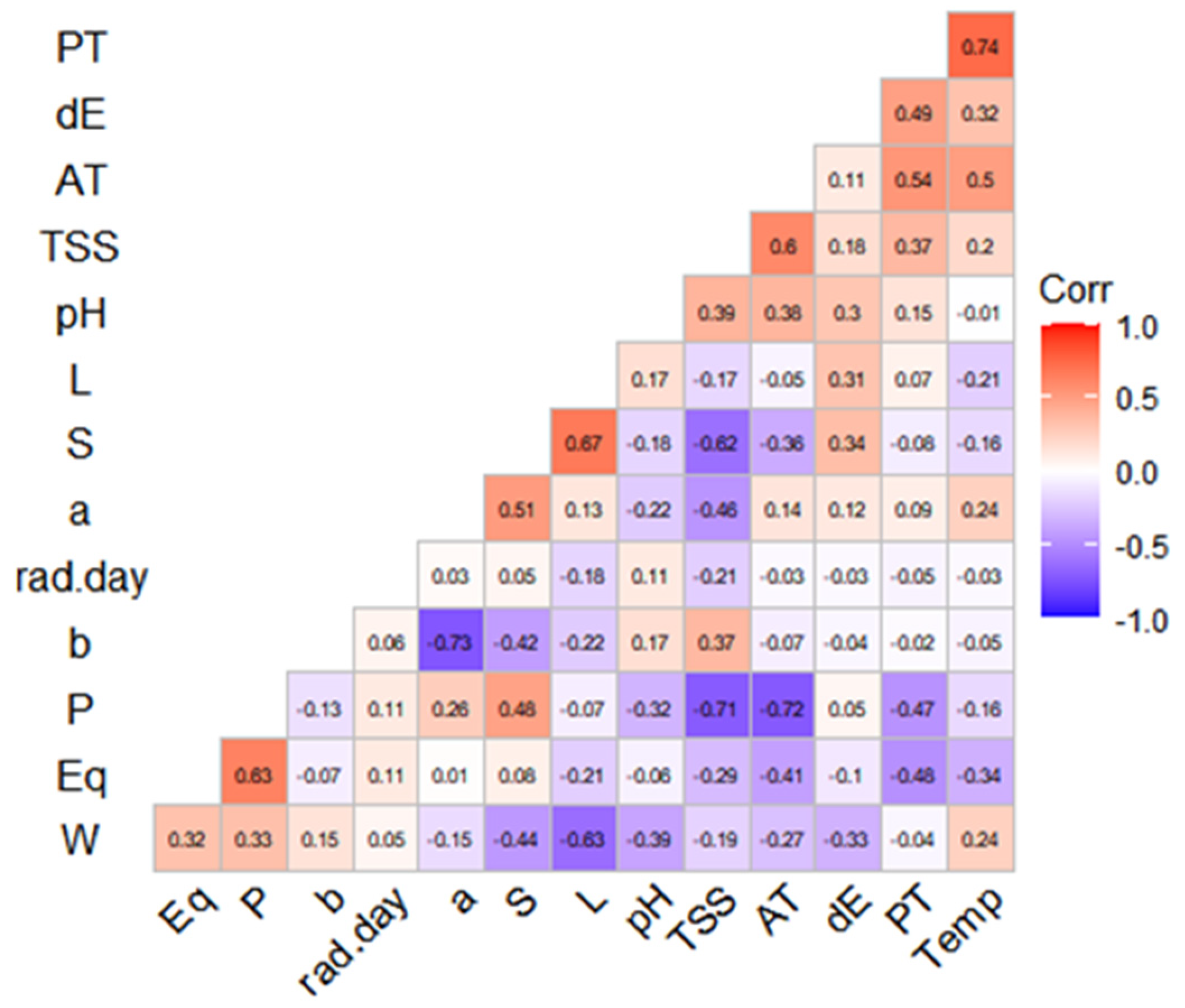
Sixteen RNA viruses that pose a significant risk for transfusion–transmission were selected for inclusion on the pathogen chip. We describe the early stage development and validation of a microarray-based platform (pathogen chip) for simultaneous molecular detection of transfusion–transmitted RNA viruses. A screening platform that could simultaneously detect all known transfusion–transmitted pathogens and allow rapid addition of new targets would significantly increase blood safety and could improve the response to new agents. Blood donors are currently screened for less than half of known agents, primarily by individual tests. New and emerging transfusion–transmitted infections remain a threat to the blood supply.
